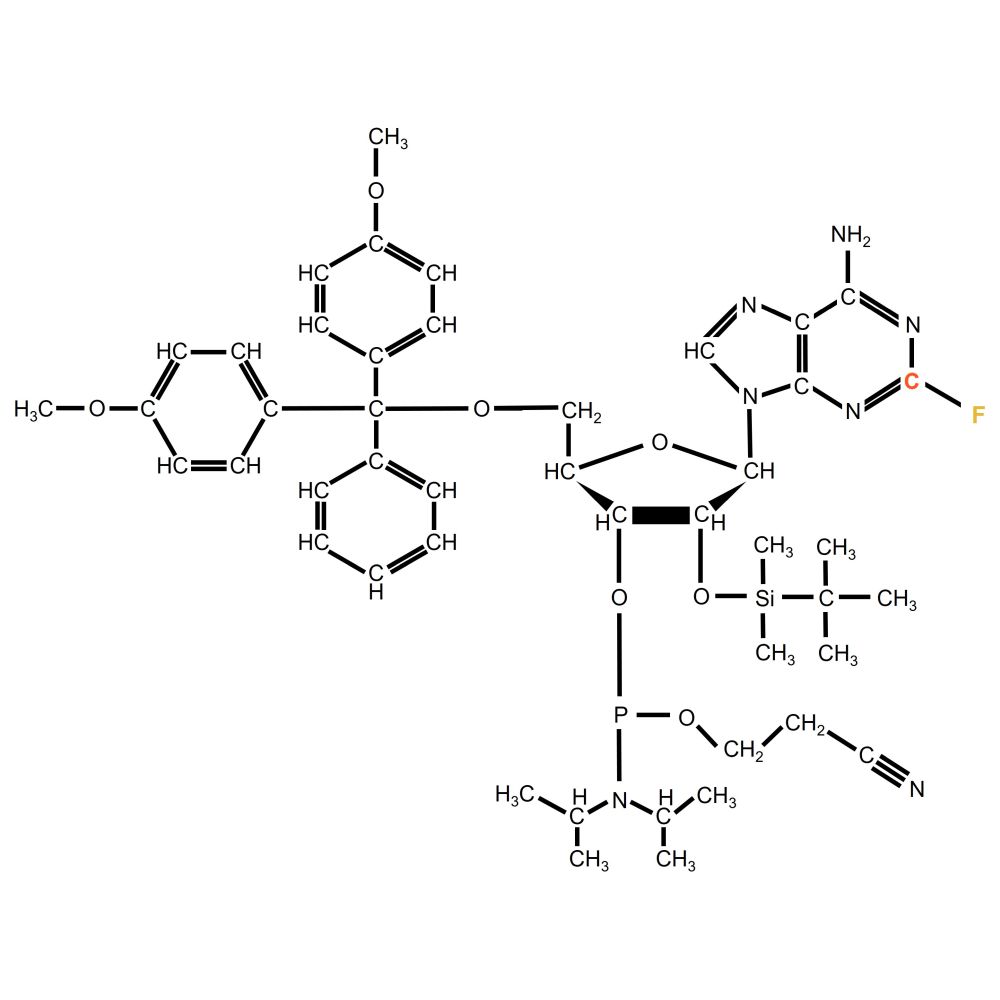
Fluoroadenosine Phosphoramidite
From: 1.170 € plus VAT, plus delivery
Synonyms: 5′-O-DMT-2′-O-TBDMS-2-Fluoradenosine-3′-CE phosphoramidite, DMT-2’O-TBDMS-2-FA Amidite
Available in various isotopic labelings and/or quantities.
Description
Detailed notation: 5′-O-(4,4′-dimethoxytrityl)-2′-O-(tert-butyldimethylsilyl)-2-Fluoradenosine-3′-[(2-cyanoethyl)-(N,N-diisopropyl)]-phosphoramidite
– This product is in powder form
– Isotopic enrichment > 98 atom %
– Chemical purity > 95 %
Packed in ND 20 crimp neck bottles. Configured for ABI.
References
| Silantes NTPs in scientific publications: | • Becette, O., Olenginski, L. T., & Dayie, T. K. (2019). Solid-Phase chemical synthesis of stable Isotope-Labeled RNA to aid structure and dynamics studies by NMR spectroscopy. Molecules, 24(19), 3476. https://doi.org/10.3390/molecules24193476 •Štih, V., Amenitsch, H., Plavec, J., & Podbevšek, P. (2023). Spatial arrangement of functional domains in OxyS stress response sRNA. RNA, 29(10), 1520–1534. https://doi.org/10.1261/rna.079618.123 |
|---|---|
| Silantes phosphoramidites in scientific publications: | • Becette, O., Olenginski, L. T., & Dayie, T. K. (2019). Solid-Phase chemical synthesis of stable Isotope-Labeled RNA to aid structure and dynamics studies by NMR spectroscopy. Molecules, 24(19), 3476. https://doi.org/10.3390/molecules24193476 •Štih, V., Amenitsch, H., Plavec, J., & Podbevšek, P. (2023). Spatial arrangement of functional domains in OxyS stress response sRNA. RNA, 29(10), 1520–1534. https://doi.org/10.1261/rna.079618.123 |
| Silantes oligonucleotide synthesis service in scientific publications: | • Belfetmi, A., Zargarian, L., Tisné, C., Sleiman, D., Morellet, N., Lescop, E., Maskri, O., René, B., Mély, Y., Fosse, P., & Mauffret, O. (2016). Insights into the mechanisms of RNA secondary structure destabilization by the HIV-1 nucleocapsid protein. RNA, 22(4), 506–517. https://doi.org/10.1261/rna.054445.115 • Borggräfe, J., Victor, J., Rosenbach, H., Viegas, A., Gertzen, C. G. W., Wuebben, C., … Etzkorn, M. (2021). Time-resolved structural analysis of an RNA-cleaving DNA catalyst. Nature, 601(7891), 144–149. https://doi.org/10.1038/s41586-021-04225-4 • Chernatynskaya, A. V., Deleeuw, L., Trent, J. O., Brown, T., & Lane, A. N. (2009). Structural analysis of the DNA target site and its interaction with Mbp1. Organic & Biomolecular Chemistry, 7(23), 4981. https://doi.org/10.1039/b912309a • Van Melckebeke, H., Devany, M., Di Primo, C., Beaurain, F., Toulmé, J., Bryce, D. L., & Boisbouvier, J. (2008). Liquid-crystal NMR structure of HIV TAR RNA bound to its SELEX RNA aptamer reveals the origins of the high stability of the complex. Proceedings of the National Academy of Sciences, 105(27), 9210–9215. https://doi.org/10.1073/pnas.0712121105 |
| Silantes 14-mer RNA Standard in scientific publications: | • Duchardt, E., & Schwalbe, H. (2005). Residue Specific Ribose and Nucleobase Dynamics of the cUUCGg RNA Tetraloop Motif by MNMR 13C Relaxation. Journal of Biomolecular NMR, 32(4), 295–308. https://doi.org/10.1007/s10858-005-0659-x • Hartlmüller, C., Günther, J. C., Wolter, A. C., Wöhnert, J., Sattler, M., & Madl, T. (2017). RNA structure refinement using NMR solvent accessibility data. Scientific Reports, 7(1). https://doi.org/10.1038/s41598-017-05821-z • Nozinovic, S., Fürtig, B., Jonker, H. R. A., Richter, C., & Schwalbe, H. (2009). High-resolution NMR structure of an RNA model system: the 14-mer cUUCGg tetraloop hairpin RNA. Nucleic Acids Research, 38(2), 683–694. https://doi.org/10.1093/nar/gkp956 • Richter, C., Kovacs, H., Buck, J., Wacker, A., Fürtig, B., Bermel, W., & Schwalbe, H. (2010). 13C-direct detected NMR experiments for the sequential J-based resonance assignment of RNA oligonucleotides. Journal of Biomolecular NMR, 47(4), 259–269. https://doi.org/10.1007/s10858-010-9429-5 • Ferner, J., Villa, A., Duchardt, E., Widjajakusuma, E., Wöhnert, J., Stock, G., & Schwalbe, H. (2008). NMR and MD studies of the temperature-dependent dynamics of RNA YNMG-tetraloops. Nucleic Acids Research, 36(6), 1928–1940. https://doi.org/10.1093/nar/gkm1183 |
| Relevant blog articles: | • What Are Stable-Isotope Labeled Nucleic Acids?: https://www.silantes.com/stable-isotope-labeled-nucleic-acids/ • Synthesizing Stable Isotope-Labeled Nucleic Acids: https://www.silantes.com/synthesizing-stable-isotope-labeled-nucleic-acids/ • The Advantages of Using Stable Isotope-Labeled Nucleic Acids: https://www.silantes.com/advantages-stable-isotope-labeled-nucleic-acids/ • Applications of Stable Isotope-Labeled Molecules: Exploring the Power of Isotopic Tracers: https://www.silantes.com/applications-stable-isotope-labeled-molecules/ • Custom RNA & DNA Synthesis Services: Tailored Solutions for Your Nucleic Acid Needs: https://www.silantes.com/custom-dna-rna-synthesis/ |
| Relevant webinars: | • Easy 13C, 15N-Labeling of DNA through isothermal amplification: Applications to G-quadruplex, aptamer, and DNAzyme: https://www.youtube.com/watch?v=HDOHcVhjZro |