Fluoro NTPs and Fluoro Phosphoramidites
Fluorine labeling is a versatile and powerful tool for NMR structural investigation of nucleic acids. The fluorine nucleus offers several attractive properties for NMR spectroscopy: high NMR sensitivity (0.83 of 1H), 100% natural abundance, and a chemical shift that responds to its local chemical environment. In addition, fluorine’s virtual absence in biological systems enables measurements that are free of background signals making it an ideal candidate for in vivo NMR applications.
(H2) 19F-13C Labeling Results in Very Favourable Spectroscopic Behaviour
Recently, Haribabu Arthanari and co-workers showed that an aromatic labeling scheme with a [19F-13C] spin pair has a very favorable spectroscopic behavior in an NMR-TROSY experiment alleviating the size problem in nucleic acid NMR.
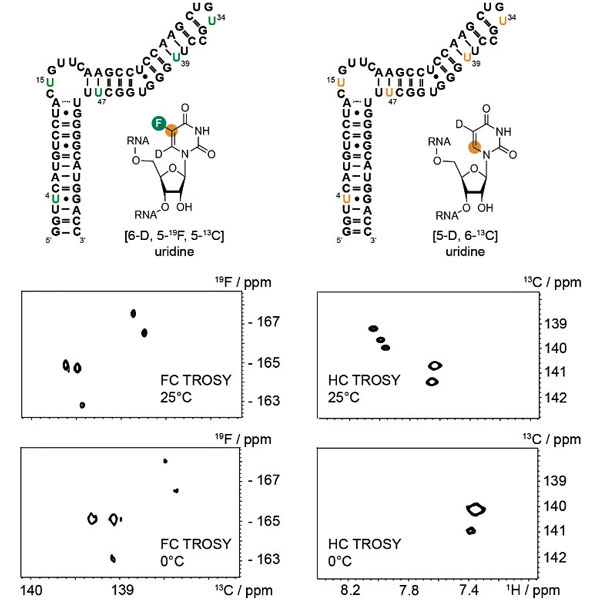
The favorable spectral behaviour of labeled fluorine presents new opportunities for characterizing the structure, dynamics, and interactions of nucleic acids with the help of appropriate fluorine NMR hardware.
[1] Nußbaumer, Felix, Raphael Plangger, Manuel Roeck, and Christoph Kreutz. “Aromatic 19F–13C TROSY—[19F, 13C]‐Pyrimidine Labeling for NMR Spectroscopy of RNA.” Angewandte Chemie (International Ed.) 59.39 (2020): 17062-7069.
References
Use cases of the Silantes NTPs in scientific publications:
- Mieczkowski, M., Steinmetzger, C., Bessi, I., Lenz, A., Schmiedel, A., Holzapfel, M., Lambert, C., Pena, V., & Höbartner, C. (2021). Large Stokes shift fluorescence activation in an RNA aptamer by intermolecular proton transfer to guanine. Nature Communications, 12(1). https://doi.org/10.1038/s41467-021-23932-0
- Musheev, M. U., Schomacher, L., Basu, A., Han, D., Krebs, L., Scholz, C., & Niehrs, C. (2022). Mammalian N1-adenosine PARylation is a reversible DNA modification. Nature Communications, 13(1). https://doi.org/10.1038/s41467-022-33731-w
- Xu, Y., McSally, J., Andricioaei, I., & Al-Hashimi, H. M. (2018). Modulation of Hoogsteen dynamics on DNA recognition. Nature Communications, 9(1). https://doi.org/10.1038/s41467-018-03516-1
- Li, M., Wang, Y., Wei, X., Cai, W., Wu, J., Zhu, M., Wang, Y., Liu, Y., Xiong, J., Qu, Q., Chen, Y., Tian, X., Yao, L., Xie, R., Li, X., Chen, S., Huang, X., Zhang, C., Xie, C., . . . Lin, S. (2024). AMPK targets PDZD8 to trigger carbon source shift from glucose to glutamine. Cell Research. https://doi.org/10.1038/s41422-024-00985-6
- Cromsigt, J., Schleucher, J., Gustafsson, T., Kihlberg, J., & Wijmenga, S. (2002). Preparation of partially 2H/13C-labelled RNA for NMR studies. Stereo-specific deuteration of the H5’’ in nucleotides. Nucleic Acids Research, 30(7), 1639–1645. https://doi.org/10.1093/nar/30.7.1639
- Rangadurai, A., Szymanski, E. S., Kimsey, I., Shi, H., & Al-Hashimi, H. M. (2020). Probing conformational transitions towards mutagenic Watson–Crick-like G·T mismatches using off-resonance sugar carbon R1ρ relaxation dispersion. Journal of Biomolecular NMR, 74(8–9), 457–471. https://doi.org/10.1007/s10858-020-00337-7
- Noeske, J., Richter, C., Grundl, M. A., Nasiri, H. R., Schwalbe, H., & Wöhnert, J. (2005). An intermolecular base triple as the basis of ligand specificity and affinity in the guanine- and adenine-sensing riboswitch RNAs. Proceedings of the National Academy of Sciences, 102(5), 1372–1377. https://doi.org/10.1073/pnas.0406347102
- Ohira, T., Minowa, K., Sugiyama, K., Yamashita, S., Sakaguchi, Y., Miyauchi, K., Noguchi, R., Kaneko, A., Orita, I., Fukui, T., Tomita, K., & Suzuki, T. (2022). Reversible RNA phosphorylation stabilizes tRNA for cellular thermotolerance. Nature, 605(7909), 372–379. https://doi.org/10.1038/s41586-022-04677-2
- Vögele, J., Duchardt-Ferner, E., Bains, J. K., Knezic, B., Wacker, A., Sich, C., Weigand, J. E., Šponer, J., Schwalbe, H., Krepl, M., & Wöhnert, J. (2024). Structure of an internal loop motif with three consecutive U•U mismatches from stem–loop 1 in the 3′-UTR of the SARS-CoV-2 genomic RNA. Nucleic Acids Research, 52(11), 6687–6706. https://doi.org/10.1093/nar/gkae349
- Broft, P., Rosenkranz, R. R., Schleiff, E., Hengesbach, M., & Schwalbe, H. (2022). Structural analysis of temperature-dependent alternative splicing of HsfA2 pre-mRNA from tomato plants. RNA Biology, 19(1), 266–278. https://doi.org/10.1080/15476286.2021.2024034
Use cases of the Silantes phosphoramidites in scientific publications:
- Becette, O., Olenginski, L. T., & Dayie, T. K. (2019). Solid-Phase chemical synthesis of stable Isotope-Labeled RNA to aid structure and dynamics studies by NMR spectroscopy. Molecules, 24(19), 3476. https://doi.org/10.3390/molecules24193476
- Štih, V., Amenitsch, H., Plavec, J., & Podbevšek, P. (2023). Spatial arrangement of functional domains in OxyS stress response sRNA. RNA, 29(10), 1520–1534. https://doi.org/10.1261/rna.079618.123
Use cases of the Silantes oligonucleotide synthesis service in scientific publications:
- Belfetmi, A., Zargarian, L., Tisné, C., Sleiman, D., Morellet, N., Lescop, E., Maskri, O., René, B., Mély, Y., Fosse, P., & Mauffret, O. (2016). Insights into the mechanisms of RNA secondary structure destabilization by the HIV-1 nucleocapsid protein. RNA, 22(4), 506–517. https://doi.org/10.1261/rna.054445.115
- Borggräfe, J., Victor, J., Rosenbach, H., Viegas, A., Gertzen, C. G. W., Wuebben, C., … Etzkorn, M. (2021). Time-resolved structural analysis of an RNA-cleaving DNA catalyst. Nature, 601(7891), 144–149. https://doi.org/10.1038/s41586-021-04225-4
- Chernatynskaya, A. V., Deleeuw, L., Trent, J. O., Brown, T., & Lane, A. N. (2009). Structural analysis of the DNA target site and its interaction with Mbp1. Organic & Biomolecular Chemistry, 7(23), 4981. https://doi.org/10.1039/b912309a
- Van Melckebeke, H., Devany, M., Di Primo, C., Beaurain, F., Toulmé, J., Bryce, D. L., & Boisbouvier, J. (2008). Liquid-crystal NMR structure of HIV TAR RNA bound to its SELEX RNA aptamer reveals the origins of the high stability of the complex. Proceedings of the National Academy of Sciences, 105(27), 9210–9215. https://doi.org/10.1073/pnas.0712121105
Use cases of the Silantes 14-mer RNA Standard in scientific publications:
- Duchardt, E., & Schwalbe, H. (2005). Residue Specific Ribose and Nucleobase Dynamics of the cUUCGg RNA Tetraloop Motif by MNMR 13C Relaxation. Journal of Biomolecular NMR, 32(4), 295–308. https://doi.org/10.1007/s10858-005-0659-x
- Hartlmüller, C., Günther, J. C., Wolter, A. C., Wöhnert, J., Sattler, M., & Madl, T. (2017). RNA structure refinement using NMR solvent accessibility data. Scientific Reports, 7(1). https://doi.org/10.1038/s41598-017-05821-z
- Nozinovic, S., Fürtig, B., Jonker, H. R. A., Richter, C., & Schwalbe, H. (2009). High-resolution NMR structure of an RNA model system: the 14-mer cUUCGg tetraloop hairpin RNA. Nucleic Acids Research, 38(2), 683–694. https://doi.org/10.1093/nar/gkp956
- Richter, C., Kovacs, H., Buck, J., Wacker, A., Fürtig, B., Bermel, W., & Schwalbe, H. (2010). 13C-direct detected NMR experiments for the sequential J-based resonance assignment of RNA oligonucleotides. Journal of Biomolecular NMR, 47(4), 259–269. https://doi.org/10.1007/s10858-010-9429-5
- Ferner, J., Villa, A., Duchardt, E., Widjajakusuma, E., Wöhnert, J., Stock, G., & Schwalbe, H. (2008). NMR and MD studies of the temperature-dependent dynamics of RNA YNMG-tetraloops. Nucleic Acids Research, 36(6), 1928–1940. https://doi.org/10.1093/nar/gkm1183
Relevant blog articles:
- What Are Stable-Isotope Labeled Nucleic Acids?
- Synthesizing Stable Isotope-Labeled Nucleic Acids
- The Advantages of Using Stable Isotope-Labeled Nucleic Acids
- Applications of Stable Isotope-Labeled Molecules: Exploring the Power of Isotopic Tracers
- Custom RNA & DNA Synthesis Services : Tailored Solutions for Your Nucleic Acid Needs
Relevant webinars:
Products:
-

5-19F Fluorocytidine 5′-triphosphate
Synonym: Fluoro rCTP Li2-salt
1.170 € plus VAT, plus delivery Add to cart
Quantity: 25mg -

Fluoroadenosine 5′-triphosphate
Synonym: Fluoro rATP Li2-salt
From: 1.140 € plus VAT, plus delivery Select options This product has multiple variants. The options may be chosen on the product page
Available in various isotopic labelings and/or quantities. -

Fluoroadenosine Phosphoramidite
Synonyms: 5'-O-DMT-2'-O-TBDMS-2-Fluoradenosine-3'-CE phosphoramidite, DMT-2'O-TBDMS-2-FA Amidite
From: 1.170 € plus VAT, plus delivery Select options This product has multiple variants. The options may be chosen on the product page
Available in various isotopic labelings and/or quantities. -

Fluorouridine 5′-triphosphate
Synonym: Fluoro rUTP Li2-salt
From: 1.140 € plus VAT, plus delivery Select options This product has multiple variants. The options may be chosen on the product page
Available in various isotopic labelings and/or quantities. -

Fluorouridine Phosphoramidite
Synonyms: 5'-O-DMT-2'-O-TBDMS-5-Fluoruridine-3'-CE phosphoramidite, DMT-2'-O-TBDMS-5FU-FA Amidite
From: 1.260 € plus VAT, plus delivery Select options This product has multiple variants. The options may be chosen on the product page
Available in various isotopic labelings and/or quantities.
